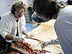
Skeleton shrimp, Caprella bathytatos, collected from the mouthparts of the deep sea spider crab. The crab was collected on Alvin dive 3807 at Warwick Seamount. Click image for larger view.
Living in the Jaws of Death
July 12, 2002
Thomas C. Shirley, Zac Hoyt, and Maggie Sexton
University of Alaska Fairbanks
Bradley Stevens and Taylor Heyl
National Marine Fisheries Service
How would you like to live in the jaws of a monster? The skeleton shrimp (Caprella bathytatos) has made a comfortable life doing just that—living on the jaws of a deep sea spider crab (Macroregonia macrocheira) that is more than 1,000 times larger than itself. The diminutive skeleton shrimp is not really a shrimp, but rather a kind of amphipod belonging to the family Caprellidae. The skeleton shrimp has several adaptations that allow it to live in this precarious location. Some of its appendages are modified to cling to the bristles found on the crab jaws. Also, the caprellid has a special pouch called a marsupium used for brooding its young; the young are thus living on a suitable host when they leave the care of their mother.
Why would a small animal choose such a precarious location to live? One reason might be that the caprellid is guaranteed a ready food source. As long as the crab has food to eat, the caprellid does too, even in a food-scarce environment like the deep sea. Another possible reason is that the location might provide safety from other predators, such as fish. The devil you know might be better than the devil you don't!
Genomics in Deep-sea Microbiology: Who is out there and what are they doing?

Naomi Ward is emptying seawater collected in Niskin bottles from the CTD. Click image for larger view.
Naomi Ward, Collaborative Investigator
The Institute for Genomic Research
What is genomics?
Genomics is the study of an organism's genome, or genetic material. The genome contains the coded instructions needed to build and maintain the organism. In most organisms, the genome consists of deoxyribonucleic acid—DNA. DNA is composed of four building blocks (the four nucleotides, also known as “bases”). The particular order of these building blocks in a string of DNA is a code for production of proteins. Proteins are key structural components of the cell and also act as enzymes—chemical catalysts for numerous processes that are essential to the cell. Genomic technologies can help us determine the identity and activities of bacteria living in diverse environments, including deep-sea habitats such as those visited during the current expedition.
Marine bacteria: key players in marine processes
Bacteria are abundant and important members of the marine environment. In a drop of seawater, there can be anywhere from a million to a billion bacterial cells. Similarly, high numbers of bacteria are found in marine sediments, even far beneath the ocean floor. These microbes have important roles in the cycling of essential elements such as carbon, nitrogen, phosphorus, and sulfur. They serve as a food source for other forms of marine life, and participate in the decay process that regenerates nutrients from dead organisms. Recent genomic studies have shed light on the structure and function of these marine bacterial communities, and have even revealed the occurrence of a new type of bacterial photosynthesis in the oceans.

The basket attached to the front of the Alvin is filled with sample collection devices such as pushcores, which are attached to the front of the basket. Alvin pilot Bruce Strickrott is detaching several Niskin bottles after the second dive on Warwick Seamount. Click image for larger view.
Sampling the seamounts
During our expedition, we have been sampling marine bacteria from deep water, surface water, deep sediments, and biofilms from the exterior surfaces of animals. Deep water samples are collected using Alvin, with a set of five bottles that are triggered by the sub pilot to collect water at different sites. An alternative device for deep-water collection is the CTD (Conductivity, Temperature, Depth), which carries 23 larger bottles that are triggered to collect at different depths. When the water samples are brought on board, they are passed through a series of filters to trap microbes of different sizes.
Deep-sea sediment samples are captured by devices called “pushcorers” that are attached to Alvin. Pushcores are hollow tubes that are pushed into the mud. Back on the ship, the pushcores can be sliced up, so that we have separate samples from different depths of sediment.
We have also been collecting some biofilm material from the outside of animals. A biofilm is a layer of bacterial cells held together on a solid surface that can be either living (e.g., a crab) or nonliving (e.g., a rock). The “glue” that holds the biofilm together is made of mucus-like substances produced by the bacteria. Biofilms are found everywhere, in both natural and human-made settings. They can pose a threat to human health when they form on indwelling medical devices such as catheters. In contrast, the biofilms that form on deep-sea surfaces are interesting because they may help other (larger) organisms such as larvae attach to and colonize the surface.

Naomi Ward collects some of the thick mucus that is oozing from this deep-sea coral after a very successful dive at Warwick Seamount. Click image for larger view.
Back at the lab
After the cruise, DNA will be extracted and purified from the samples. To determine which bacteria are present, we will sequence a particular gene (the 16S ribosomal RNA (16S rRNA) gene) that all bacteria possess and that we can use as a “marker” gene. If there are 500 different bacteria in a given sample, we will get 500 different 16S rRNA gene sequences. We can identify these by comparing them to a database containing all known sequences from this gene. To determine what these bacteria are doing in their habitat, we will sequence larger pieces of DNA, using the same methods that we apply to sequencing whole genomes of cultured bacteria. Comparison of these sequences to the genomic databases will allow us to assign a tentative function to each sequence.
Applications
The ultimate goal of applying genomic approaches to studying deep-sea bacterial communities is to identify which bacteria are present and what they are doing. If we can determine which proteins are being produced, we can reconstruct the pathways of metabolism for these different bacteria. We will know what substrates (food-stuffs) they use for nutrition, and how they bring them into their cell and break them down. We may also catch a glimpse of how bacteria fight each other and defend their territory, through information from genes that encode antibiotics and other substances.
Genomic information captured from bacterial communities may also reveal the presence of bacteriophages. These viruses, which prey on bacteria, often insert their own DNA into the bacterium's DNA, as part of their sneaky strategy to take over the bacterium's replication machinery and make thousands of copies of themselves. Bacteriophages are thought to be one of the key forces in shaping the structure of natural bacterial communities. Additional information on their distribution and association with particular bacteria may help us understand this process.
Sign up for the Ocean Explorer E-mail Update List.