
By Susie Cummings, Graduate Student, Oregon State University
September 28, 2020
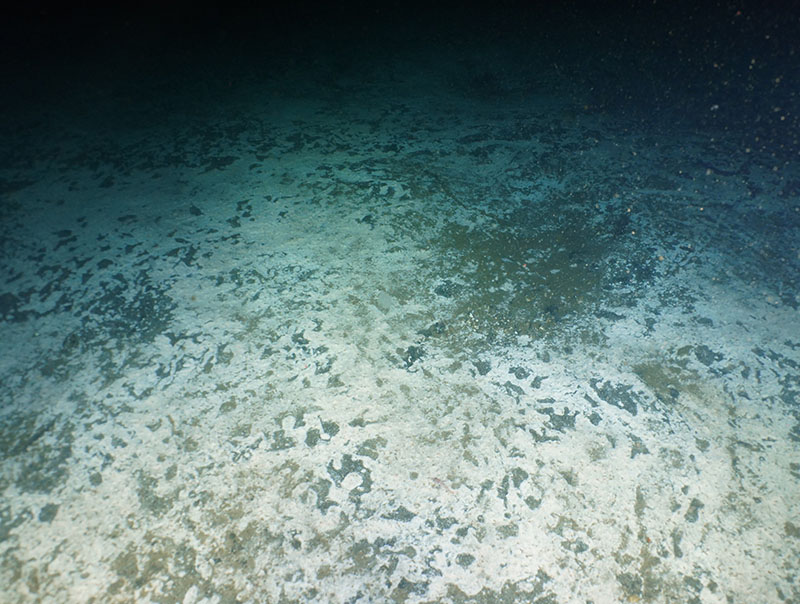

While microbial mats at methane seeps, like this one, are home to many microbes, deep-sea sediment has a billion bacteria in every milliliter of mud, whether at a seep or in the middle of the ocean. Image courtesy of A. Thurber, Oregon State University, OCNMS, NOAA OER, and Ocean Exploration Trust: NA-121. Download larger version (jpg, 2.8 MB).
How would you conduct a census for bacteria?
As scientists, we want to learn about the diversity of different life on Earth – and that includes microscopic life as well! But how do you get information about things so small they can’t be seen with the naked eye?
Imagine if we could conduct a census for microscopic life. We might ask the microbes questions about who they are and what kind of jobs they do. We want to learn more about their lives, so we might also ask them what kind of foods they eat, who they interact with, and what sort of things they can make. The answers to these questions would give us a picture of the microbial community in a particular area. And like different cities and towns of people, each microbial community is different based on where they are, who is there, and what is happening there.
While scientists cannot literally go door-to-door to collect information on microbes, their research can be thought of as a kind of census. Image courtesy of Susie Cummings, graduate student at Oregon State University. Download larger version (jpg, 1 MB).
This “Microbial Census” is not just imaginary! We can already conduct these kinds of surveys using a tool called “metagenomics.” A genome is the biological blueprint contained within each living thing, and genome sequencing is a method of reading these blueprints. When we read (or sequence) each genome, we learn information about who the organism is: how it is shaped, what kinds of things it can “sense,” what foods it can eat, and what methods it uses to communicate with others. When we sequence a combined collection of different genomes – from a whole “city” of microbes, for instance – that is a metagenome. In most cases, these metagenomes are really a jumble of different small pieces of genomes, and so we must use computing methods to help us sort through and identify these various pieces.

The Omics Toolbox contains various tools used to measure a community, including metagenomics, transcriptomics, proteomics, and metabolomics. Image courtesy of Susie Cummings, graduate student at Oregon State University. Download larger version (jpg, 573 KB).
There are other tools related to metagenomics, called “transcriptomics,” “proteomics,” and “metabolomics” (among others), and the term “omics” is often used to refer to this whole set of tools. Each of these tools gives us slightly different information. Metagenomics focuses on genomes, which are the overall “templates” for organisms, but this does not tell us what each organism is doing in the moment. For that purpose, transcriptomics, proteomics, and metabolomics are used; they measure different chemical messengers to give us a better sense of what is currently happening in that community. These answer questions such as: How much activity is occurring? What products are being made? What messages are being passed between different organisms?
When we combine different omics methods at once, we get a more holistic picture of the community, and can even observe how microbes interact with larger creatures!

“Take me down to Microbial City, where the worms are red and the ground is gritty.” Image courtesy of Susie Cummings, graduate student at Oregon State University. Download larger version (jpg, 1 MB).
So, why are we using omics to study the deep sea? We want to answer questions about how food and energy flows through the ocean, and omics can help track these flows. We want to discover potential novel medicines, and omics can reveal how they are made, so that we can recreate them in the laboratory. Finally, we want to figure out how life survives in the numerous unusual habitats on Earth, and omics helps us describe this very cool “microbial city”!